 |
de | fr | en Druckansicht ![]()
3R-INFO-BULLETIN 36
January 2008
Author

Professor Pierre Cosson's research group (including Mohammed Benghezal, Laeticia Alibaud, Romain Froquet) has been using Dictyostelium as a model to study host-pathogen interactions for the past 10 years. Collaboration on the part of several researchers has allowed the dissemination of these techniques to other laboratories and the creation of the NEMO network (project no. 99-05), dedicated to the development of alternative models to study bacterial pathogens.
Current address:
Prof. Dr. Pierre Cosson
pierre.cosson@medecine.unige.ch
Centre Médical Universitaire, Geneva Faculty of Medicine
1 rue Michel Servet
CH-1211 Geneva 4
Switzerland
Editor
Peter Maier, Scientific Adviser of the 3R Research Foundation
Host pathogen interactions can be studied in amoebae instead of animals
Studying the interaction of pathogenic bacteria with a host[*] necessitates the controlled infection of a host. Mice or rats are classical models and these experiments inflict significant suffering on infected animals. In the present project no. 90-03, supported by the 3R Research Foundation, unicellular Dictyostelium amoebae were used as alternative hosts to study bacterial pathogens. The results obtained demonstrate that this model could, to a large extent, replace the use of mammals.
Amoebae against bacteria
After a pathogen breaches the first physical barriers (e.g. damaged skin) of the host, phagocytes represent an important part of the constitutive (in contrast to the immunological) cellular defence.
Amoebae are complex unicellular organisms that have been used largely for genetic studies. They engulf and kill bacteria in a manner similar to mammalian phagocytes (Fig. 1). Therefore they can be used as a model for phagocytic mammalian cells. On the other hand, bacteria have developed a wide variety of strategies to avoid phagocytosis and killing by using similar mechanisms to fight amoebae or mammalian phagocytes. They can prevent growth of amoebae by avoiding phagocytosis, by secreting a wide variety of toxic compounds, or due to their resistance to intracellular killing. All these bacterial virulence traits are essential during the infection of a mammalian host, and all of them can be measured by using a Dictyostelium host.
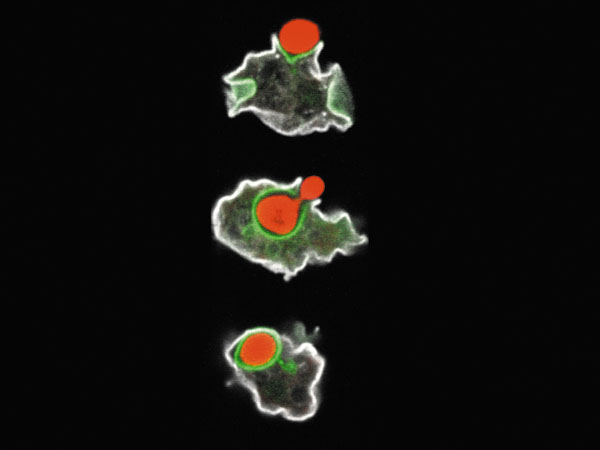
Fig. 1: An amoeba (white) eating a yeast (red)
Virulence of Pseudomonas bacteria in mammals and amoebae
Pseudomonas bacteria represent a serious health problem in developed countries and are responsible notably for pneumonia or septicemia. They are also often resistant to a wide spectrum of antibiotics. Classical models to assess the virulence of Pseudomonas bacteria include burn wound infections or pulmonary infections, usually in immuno-compromised mice or rats. Our initial studies demonstrated that Dictyostelium amoebae can grow on non-virulent Pseudomonas strains, but not on virulent strains1 (Fig. 2). In a more recent series of experiments, we mutagenized randomly virulent Pseudomonas bacteria, isolated mutants that were non-virulent against Dictyostelium, and tested them in a classical mouse pulmonary infection model2. The mutants all showed a strong reduction in their virulence against mice. Together these results indicate that the Dictyostelium model system can be used as a non-mammalian model to assess Pseudomonas virulence.
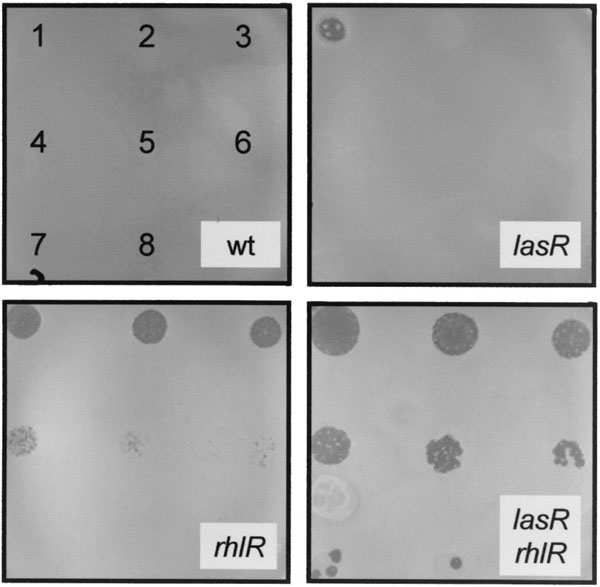

Fig. 2: Quantitative assessment of Pseudomonas virulence against Dictyostelium discoideum. Dictyostelium cells were applied as droplets onto a lawn of P. aeruginosa bacteria, as represented in the upper left panel. The numbers of Dictyostelium cells applied were 50,000 in drop number 1, 10,000 in 2, 2000 in 3, etc. Virulent wild-type bacteria (wt) inhibited growth of Dictyostelium, rhlR and lasR mutations rendered bacteria permissive for Dictyostelium growth.
Assessing fish pathogens
In an effort to extend our results to other bacterial pathogens, we defined conditions in which the virulence of various Aeromonas species could be tested against Dictyostelium. Aeromonas salmonicida infects cold-blooded vertebrates and therefore mammals are not relevant hosts. This necessitated an adjustment of the assay conditions. Again Dictyostelium predicted correctly the virulence of Aeromonas salmonicida, but also of Aeromonas hydrophila, which can also be pathogenic for humans 3.
Towards an extensive analysis of host-pathogen interactions
One critical parameter determining the outcome of a bacterial infection is the potency of the host defences. Thus immuno-compromised patients can be infected by bacterial strains that would usually not infect healthy individuals. Numerous laboratories are analyzing which host genes are essential for the immune defence against various pathogens. A large number of these experiments rely on the use of transgenic mice affecting the function of various host genes.
We are using the Dictyostelium model to analyze host genes implicated in the resistance to bacterial pathogens. Our recent studies revealed the role of two amoebae proteins (Phg1 and Kil1) in the intracellular killing of internalized bacteria by Dictyostelium amoebae (Fig. 3)4. In this system, bacterial genetics and host genetics can be combined to analyze efficiently the complex confrontation of host cells with pathogenic bacteria. For example we could show that phg1 mutant amoebae are incapable of killing normal virulent Klebsiella bacteria, but they can kill Klebsiella mutants that are particularly sensitive to killing by amoebae. Phg1 is also involved in the defence of flies against bacterial infections, suggesting that the genes and mechanisms involved are most likely well conserved and relevant to the situation in mammals.
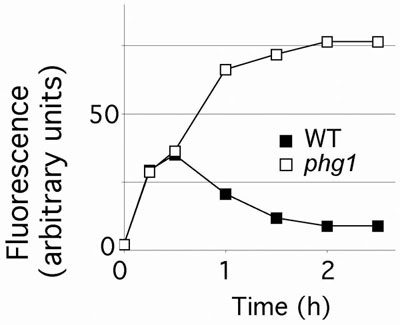
Fig. 3: Phg1 mutant Dictyostelium amoebae do not efficiently kill Klebsiella bacteria. Dictyostelium cells were incubated with Klebsiella bacteria expressing a fluorescent GFP protein. At the indicated times the intracellular fluorescence was measured. Wild-type cells internalize bacteria and then kill them, destroying the GFP fluorescence. Phg1 mutant cells internalize bacteria but fail to kill them and accumulate live fluorescent bacteria intracellularly.
3R Benefits
In order to analyze host-pathogen interactions, hosts must be infected experimentally with pathogenic bacteria. When carried out in mice or rats, this can lead to sickness or even death of the animals. These experiments are typically performed either to measure bacterial virulence or to determine the role of host genes in host-pathogen interactions. Furthermore efficient infections are often achieved in rodents by destroying some elements of their defences, for example by creating wounds or by destroying their phagocytic cells.
Clearly the present in vitro model demonstrates that bacterial virulence is similar in a Dictyostelium host model when compared to mammals. This remains true for a variety of pathogenic bacteria so far analyzed. In addition this model system can be adjusted to bacterial pathogens living in different environments. Finally our results suggest that at least a part of the analysis of host gene products involved in the defence against pathogens could be done in Dictyostelium. Our laboratory is currently actively pursuing this line of research.
In conclusion, the development of easy, cheap and reliable alternative non-mammalian models should significantly reduce the need to infect mammals. Far from hindering research in this field, we expect that use of alternative models will actually help to increase our understanding of bacterial infections.
PDF version of this Bulletin No. 36
References:
- Cosson, P., Zulianello, L., Joint-Lambert, O., Faurisson, F., Gebbie, L., Benghezal, M., van Deden, C., Kocjancic-Curty, L., Köhler, T. 2002. Pseudomonas aeruginosa virulence analyzed in a Dictyostelium discoideum host system. J. Bact. 184:3027-3033.
- Alibaud, L., Köhler, T., Coudray, A., Prigent-Combaret, C., Bergeret, E., Perrin, J., Benghezal, M., Reimmann, C., Gauthier, Y., van Delden, C., Attree, I., Fauvarque, M.O., Cosson, P. 2007. Pseudomonas aeruginosa virulence genes identified in a Dictyostelium host model. Cell. Microb. In press
- Froquet, R., Cherix, N., Burr, S., Frey, J., Vilches, S., Tomas, J.M., Cosson, P. 2007. An alternative host model to evaluate Aeromonas virulence. Appl. Environmental Microb. 73: 5657-9.
- Benghezal, M., Fauvarque, MO., Tournebize, R., Froquet, R., Marchetti, A., Bergeret, E., Lardy, B., Klein, G., Sansonetti, P., Charette, S.J., Cosson, P. 2006. Specific host genes required for the killing of Klebsiella bacteria by phagocytes. Cell. Microb. 8:139-48.
| [*] | Assessment of bacterial virulenceBacterial pathogens represent a major health problem worldwide. In developing countries they are one of the major killers (notably M. tuberculosis) particularly for immuno-compromised patients (aging people, HIV-infected patients, grafted patients) or people suffering from specific diseases (e.g. cystic fibrosis patients). One aggravating factor is the rise of new strains of bacteria resistant to one or several antibiotics, and the very limited number of newly developed antibiotics. This has led to a renewed interest in understanding how pathogenic bacteria can cause diseases. Bacteria are defined as virulent (or pathogenic) based on their ability to mount a harmful infection in a host. Virulent bacteria can infect a host, non-virulent bacteria cannot. A successful infection relies on a series of specific bacterial traits, such as the ability to secrete toxins, to escape the host immune system, to replicate within host cells or to inhibit phagocytic engulfment. For any given pathogenic bacteria, the analysis of virulence genes necessitates many experiments. Different arrays of virulence genes are involved in different bacterial species, and in different strains within the same species. Assessing bacterial virulence requires infecting a host with a potentially pathogenic bacterium and following the process of infection. The most common hosts are rodents (mice or rats), based on the notion that their immune defences are most similar to those of human patients. These experiments are particularly problematic ethically, since they inflict significant suffering on infected animals. This has led to the development of alternative hosts to follow the process of bacterial infections e.g. Drosophila flies, Caenorhabitis elegans nematodes and, as in the present project, Dictyostelium amoebae. This system is particularly simple and could in many cases replace mammalian hosts. |
| Letzte Änderung: 06.02.2008 |