 |
de | fr | en print view![]()
3R-INFO-BULLETIN 20
May 2002
The Authors
  |
The project was carried out under the guidance of Dr. F. Bootz, Head of Animal Health Services and PCR Diagnostics, Institute of Laboratory Animal Sciences, University of Zürich and Prof. Dr. F.R. Homberger, Associate Director of the Research Animal Resources Center and Head of Biosecurity and Animal Health Service, Cornell University New York. The investigations are described in three dissertations, authored by Dr. D. Popovic, Dr. M.E. Tischhauser and Dr. I.U. Sieber.
Current address:
Dr. Frank O. Bootz fobo@ltk.unizh.ch
Institute of Laboratory Animal Sciences
University of Zürich
Winterthurerstr. 190
CH-8057 Zürich
Switzerland
Prof. Dr. F. R. Homberger hombergf@mskcc.org
Memorial Sloan-Kettering Cancer Center and
Weill Medical College of Cornell University
New York, 10021
USA
Editor
Peter Maier, Scientific Adviser of the 3R Research Foundation
Animal-free screening of biological materials for contamination by rodent viruses
DNA in vitro instead of antibodies in vivo
The mouse antibody production (MAP) test[*], traditionally used to screen biological material prior to their use in vivo, has a number of drawbacks. One is the fact that animals are required for the test. Since most of the pathogens that are being screened for do not cause clinical disease in immunocompetent adult animals, the pain and distress suffered by the test animals is usually minimal. However, some infectious agents (e.g. ectromelia virus) may cause disease or even death. These cannot be ruled out in advance since the types and concentrations of viruses present in the test material are unknown. In addition, the MAP test is fairly costly and slow (5-6 weeks). There are no reliable numbers, but it is estimated that 200-300 MAP tests are conducted annually in Germany alone.
Previous attempts to replace the MAP test with an in-vitro assay using virus isolation in cell culture were unsatisfactory: not all viruses could be detected reliably in culture. With the advent of the polymerase chain reaction (PCR), a new, highly sensitive and specific tool became available. In order to replace the MAP test, it was necessary to establish PCR assays to detect all the possible viral agents being screened for (Table 1), otherwise a MAP test would still have been required to rule out the remaining pathogens.
Development of PCR Assays
For each pathogen (Figure 1), specific primers were designed based on published sequencing information. Primers were situated in conserved regions of the respective genomes to ensure that all known substrains of a virus could be detected. Where available, published PCR assays were used if they met the above criteria. All PCR assays were tested for their specificity and sensitivity using virus stock cultures at various dilutions. Virus stocks were obtained from American Type Culture Collection and various collaborators. These assays were published as part of two dissertations [Popovic (1998), Tischhhauser (1999)].
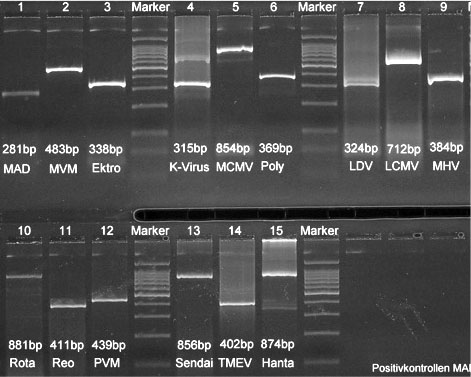
Fig. 1: Size of the PCR-Products in Clearose Gels from the different viruses tested. Abbrevations used see Table 1.
Semiquantitative analysis improves the interpretation
A TaqMan� DNA amplification system was provided by a generous grant from the Doerenkamp Zbinden Foundation. This system allowed us to follow the PCR reaction kinetics in real time (Figure 2): Each reaction well is equipped with optical fibers. Laser light transmitted through these fibers stimulates target molecules in the reaction mix to emit fluorescent light. These target molecules are present in an inactive form on the primers and are cleaved off, becoming active, when the primer anneals and starts the strand formation. Fluorescence is measured every 5 seconds; the light emitted is directly proportional to the amount of primer used and therefore to the number of new amplicons. This method allows a semi-quantitative analysis of the results as the amplification progresses, i.e. in real time. The test was named Infectious Microbe PCR Amplification Test (IMPAT).
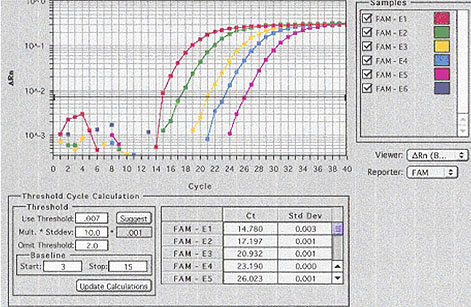
Fig. 2: Taqman� (realtime PCR). Pictorial representation of the amplification rate of the virus serial dilutions. For example, the amplification of the highest virus-concentration (red line) raised at cycle 14, the lowest concentration (purple line) at cycle 26.
IMPAT, a replacement for the MAP-Test
In order to validate the IMPAT, it was compared with the MAP test. Samples of known concentration (tissue culture infective dose) were generated for each of the 14 viruses on the screening panel (Table 1). For each virus four 100-fold serial dilutions were made.
MAP test: Groups of 2 mice were inoculated with each virus at each of the dilutions. After 30 days the animals were euthanized and exsanguinated. The serum was tested for antibodies specific to the respective viruses using immunofluoresence assays.
IMPAT: To mimic routine conditions, the virus dilutions were not analyzed directly by IMPAT, but rather used to inoculate a standard tumor cell suspension. The tumor cells were incubated with the virus samples for 1 hour (DNA viruses) or 5 minutes (RNA viruses) before DNA or RNA extraction. Nucleic acids isolated from the contaminated tumor material were analyzed using standard PCR and the TaqMan� method.
Sensitivity: The three methods were equally sensitive in detecting two of the 14 viruses (Bootz und Sieber, 2002). One of these was mouse hepatitis virus, which was lethal in 7 of 8 mice inoculated. While no serology could be done the samples were considered positive. The molecular methods were more sensitive than the MAP-Test in detecting all the other pathogens. This was the case for six of the twelve remaining viruses when using the conventional PCR and for all 12 when using the TaqMan�. The fact that real time PCR was in many cases more sensitive than the standard PCR was attributed to two reasons: 1) The TaqMan� methodology using fluorescence emission is considered to be more sensitive, and 2) a standard annealing temperature was used for the conventional PCR assays, while for the TaqMan� real time PCR the annealing temperature was optimized for each primer set.
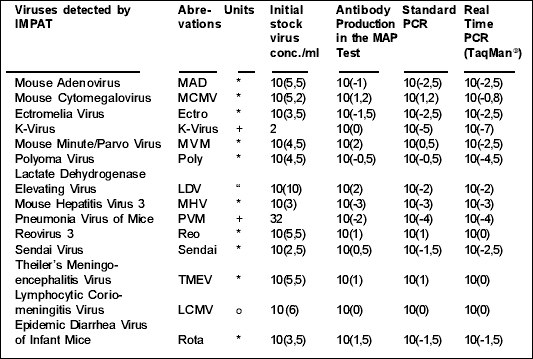
Table 1 List of viruses detected by the IMPAT and limits of detection:
*=TCID50=50% Tissue Culture Infectious Dose
+=HA-units=Haemaglutinin units
“ =ID50=Infectious Dose (50% of mice)
o=PFU=Plaque Forming Units
( ) = indices of dilution factors.
Important advantages of the in vitro method
The specificity and the sensitivity of the IMPAT is at least equivalent and in most cases superior to that of the MAP test. In addition there are a number of other advantages over the in vitro method:
- No animals are required (replacement)
- Turn-around time is reduced to 1-2 days (MAP test 5-6 weeks)
- Independent of the individual immune response of the mouse (smaller variability)
- No animals inoculated with unknown infectious agents need be housed in the animal facility (no risk to the animal facility or the investigators)
The main difference between the MAP test and IMPAT is that the MAP test detects only the immune response of a mouse to infectious virus particles whereas IMPAT detects the presence of nucleic acids from infectious but also from non-infectious, inactivated virus. Here IMPAT has a further clear advantage, since detection of inactivated virus is a warning sign that a contamination had occurred at some time and such material should be excluded due to the increased risk associated with it.
Negligible drawback
The only drawback of IMPAT is its high specificity. While serological methods are based on the immune reaction to multiple proteins, IMPAT detects only a very specific sequence of the viral genome. This carries the risk that it may not detect a mutant variant of the pathogen. We have tried to keep this risk to a minimum by carefully choosing the primers in regions of the genome that are highly conserved. The IMPAT is now available as a routine service (www.ltk.unizh.ch).
Published updated version of this Bulletin 20/2007 (PDF)
References:
- Parker, J. C. and R. K. Reynolds (1968). "Natural history of Sendai virus infection in mice." Am. J. Epidem. 88: (112-125)
- Bootz, F. und Sieber, I. (2002). "Ersatzmethode für den Maus/Ratte Antikörper Produktions Test. Sensitivitätsvergleich zwischen in vitro und in vivo Methode." ALTEX 19, Suppl. 1 (in press).
- Dissertations from the Institute of Laboratory Animal Sciences. University of Zürich:
- Popovic, D. (1998). "Die Polymerase-Kettenreaktion (PCR) als Ersatz für den Mäuse- und Ratten-Antikörper-Produktionstest."
- Tischhauser, M. E. (1999). "Replacement of mouse and rat antibody production (MAP/RAP) test by polymerase chain reaction."
- Sieber, I.U. (2001) "Ersatzmethode für den Maus-Antikörper-Produktionstest. Ein Sensitivitärsvergleich verschiedener PCR-Assays mit dem Tierversuch."
| [*] | Mouse Antibody Production Test (MAP Test)Biological materials, such as serum, transplantable tumors, cell cultures and hybridoma lines, that originate from or have been passaged through rodents may be inadvertently contaminated by rodent pathogens. If inoculated into rodents they may transmit these infectious agents to the graft recipient. The resulting, generally subclinical, infection may not only alter the outcome of the specific experiment, but may also be transmitted to other animals in the colony. It is therefore paramount that all biological materials be screened before they are inoculated into rodents. Traditionally this is carried out by means of the MAP test [Parker (1968)], which involves inoculating the test material into the relevant rodent species and serologically testing the inoculated animals 30 days later for antibodies specific to known viral pathogens. |
| Last modified 2008/01/30 |