 |
de | fr | en print view![]()
3R-Project 96-05
Assessment of pain and stress in mice by monitoring gene expression changes
Paulo Cinelli
Institute of Laboratory Animal Science, University of Zurich, 8057 Zurich, Switzerland
paolo.cinelli@ltk.uzh.ch
Keywords: mice; brain; pain / stress; microarray; molecular biology: pcr; refinement
Duration: 3 years Project Completion: 2009
Background and Aim
The overall aim of this project is to develop a rapid, reliable and objective method to assess pain and stress in mice on a molecular level. It is an extension of preliminary investigations using microarrays and complements ongoing studies at our Institute using telemetry and/or behavioural parameters to assess pain and stress in mice (see project 71-00). The problems proposed to be solved in the present study are, first of all, the identification of additional indicator genes and the validation of these genes using biopsy material from surgical models monitored by telemetry. A further step will be the development of a low-density microarray or a set of RT-PCR reactions with the relevant genes in order to monitor pain and/or stress. Finally, we would like to perform comparative molecular monitoring of surgical models and selected genetically modified mice (disease models from our breeding colonies) in order to validate the approach.
This molecular tool will be then used for the identification of different pain and stress levels in mice after experimental manipulations (e.g. surgery or pharmacological tests) and also for checking the housing conditions. In particular this tool is planned to be used to monitor potential pain levels in genetically modified mice
Method and Results
After careful research in available literature, we could identify about 300 genes related to pain, stress, and anxiety and selected 289 genes for spotting in triplicates on a low density microarray. We decided to spot 70 nucleotide long oligomers because this system allows a minimization of the secondary structure, high Tm's, and therefore a normalized hybridization temperature (Fig. 1).
We defined time points in postoperative mouse models for microarray analysis using a telemetric system. We are currently analyzing two different models, a first one where the animals are exposed to moderate pain and a second one with mild pain conditions. In order to identify the best suitable tissues we are currently analyzing expression pattern changes in total RNA pools from different tissues (e.g spinal cord, brain stem, hippocampus, cortex).
Preliminary hybridizations with triplicates were already performed with total RNAs isolated from whole brain. After a first comparison between the data obtained we could identify some genes whose expression was significantly changed in mice with pain.
At the moment, we are validating these genes with real time PCR.
Conclusions and Relevance for 3R
All experimental work with animals has to be monitored by a careful assessment and minimization of pain and stress. The same holds true for breeding of mutant animals. The microarray analysis will be used in parallel to behavioral observations in order to clearly define distress in genetically modified animals. By using post-mortem biopsy material, the microarray technology is much gentler in regards to animal welfare than other approaches like telemetry studies. Furthermore, this system will allow a clear dissociation between phenotype-linked data and artifacts due to the presence of pain/stress in the analyzed animals and therefore reduce the number of animals needed. This analysis tool will be very important for the improvement of anesthesia and analgesia in order to combine the best experimental conditions for both the animals and the needs of the experimentator, allowing a refinement of the animal experiment and at the same time a reduction of the number of animals. Last but not least we think that the objective assessment of pain/stress in mutant animals will play a crucial role in deciding which breeding strategy to choose in order to substantially minimize the number of affected mutant animals in breeding colonies.
(see also 3R-INFO-BULLETIN Nr. 39)
References
Paolo Cinelli, Andreas Rettich, Burkhardt Seifert, Kurt Bürki, Margarete Arras (2007) Comparative analysis and physiological impact of different tissue biopsy methodologies used for genotyping of laboratory mice. Lab Anim. 41 (2), 174-84.
Margarete Arras, Andreas Rettich, Paolo Cinelli, Hans P. Kasermann, Kurt Bürki (2007) Assessment of post-laparotomy pain in laboratory mice by telemetric re-cording of heart rate and heart rate variability. BMC Vet Res 2007, 3:16.
Paolo Cinelli, Margarete Arras and Kurt Bürki (2006) Detection of pain and stress by monitoring gene expression. ALTEX, 23, Spl, 416-419.
Figures

Figure 1A: Examples of signal obtained from total brain RNA reverse transcribed in presence of aminoallyl-modified dUTP and subsequently labeled with a cy-dye hydroxysuccinimide ester. Representative picture of an array-hybridization with aRNA labeled with aaUTP and hydroxysuccinimide ester of the cy-dyes (cy5= ctrl, cy3= brain of a mouse 5 days after telemetry transmitter implantation)
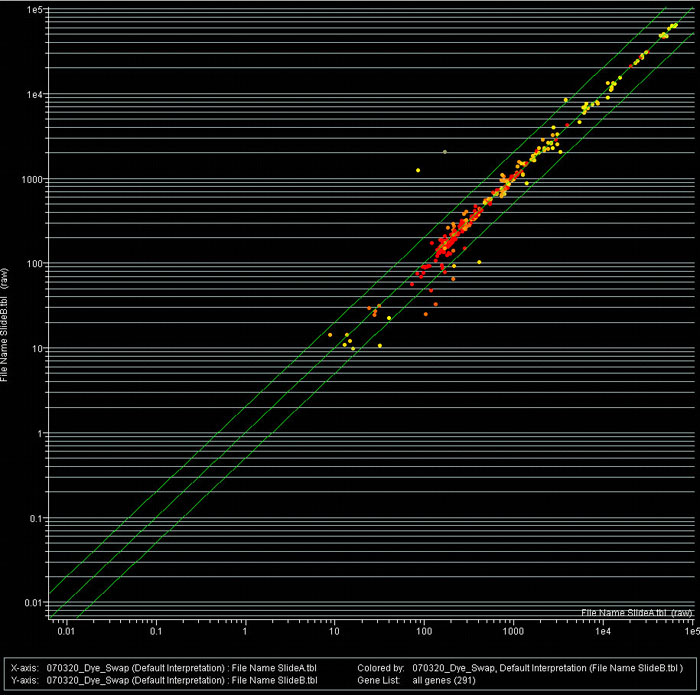
Figure 1B: Scatter plot analysis of a pain/stress microarray.
| Last modified 2018/10/12 |