 |
de | fr | en print view![]()
3R-Project 128-11
Genetic modification of the human airway epithelium – a paradigmatic system to study host responses to human respiratory viruses
Ronald Dijkman, Hulda Jonsdottir and Volker Thiel
Institute of Immunobiology, Cantonal Hospital St. Gallen, 9007 St. Gallen, Switzerland
Current address: Institute of Virology and Immunology IVI, Bern and Mittelhäusern, and Vetsuisse Faculty, University of Bern, Switzerland.
ronald.dijkman@vetsuisse.unibe.ch, hulda.jonsdottir@vetsuisse.unibe.ch, volker.thiel@vetsuisse.unibe.ch
Keywords: human; viruses; epithelia; epithelia; lung; lung diseases; cell cultures: organ-specific; cell cultures: transgenic; reduction; replacement; infectiosity
Duration: 2 years Project Completion: 2013
Background and Aim
The epithelium of the human airways is a port of entry for many respiratory viruses, such as the respiratory syncytial virus, rhinoviruses, influenza viruses, human parainfluenza viruses, adenoviruses, human metapneumoviruses and human coronaviruses. Recent advances in the cultivation of primary human airway epithelia (HAE) have permitted the study of human respiratory viral infections within a culturing system that morphologically and functionally resembles the human airways in vivo. Notwithstanding these recent technical achievements, the analysis of basic virus-host interactions at the molecular level is still preferentially performed in animal models. This is particularly true in the case of viral infections for which a murine model is available. The main reason for this preference is that when using a murine model of viral infection, advantage can be taken of widely available transgenic or knock-out mouse strains. Thus, in a murine model, and in contrast to primary human cells, individual genes can be genetically deleted or trans-complemented, in the mouse model, thereby permitting detailed analyses of virus-host interactions at the molecular level. To overcome the limitation in the human case, we endeavoured to render the HAE-culturing system amenable to genetic modifications. The research objectives of this project included: (i) the generation of lentiviral vectors encoding reporter proteins and/or shRNA-sequences that are expressed upon induction, and (ii) optimization of the transduction of primary HAE-cultures and establishment of efficacious affinity-selection methods to obtain HAE-cultures with a high level of lentiviral vector-based gene expression.
Method and Results
Construction of lentiviral vectors
To render the HAE-culturing system amenable to genetic modification, we constructed lentiviral vectors encoding the Cherry-Picker transgene and containing an insertion site for a shRNA-sequence that is under the regulation of an inducible human U6-promoter element that is actively repressed by the lactose-repressor protein. This basic organization of our lentiviral vectors has two advantages. Firstly, the Cherry-Picker gene encodes a chimeric red fluorescent protein which facilitates the detection of transduced cells. By virtue of its cell-surface expression profile and by making use of a Cherry-Picker-specific antibody, transduced cells can be sorted. Secondly, the inducibility of shRNA-expression permits a controlled knock-down of targeted genes, including those that may interfere with the differentiation of HAE-cultures.
The Cherry-Picker gene was fused via a 2A-autoprotease sequence to the LacL-gene. Unfortunately, our attempts to select the transduced cells by MACS-sorting were repeatedly unsuccessful. Further analyses revealed that the Cherry-Picker protein appeared to be retained within the cytoplasm of the transduced cells. Similar cytoplasmic retention has occasionally been described for genes that are cloned upstream of the 2A-autoprotease sequences. We have, therefore, exchanged the 2A-autoprotease sequence for an internal ribosomal entry site. Since we have already cloned shRNA-sequences that are directed against GFP and human CD13 (receptor of the human coronavirus 229E), we now proceed with the Cherry-Picker-based selection.
The overall strategy - assessing transduction efficacy - cellular composition of transduced epithelia
Lung epithelial cells were isolated from human tissue by protease digestion and were then expanded in two-dimensional cultures. The cells were either cryo-preserved or seeded in liquid-liquid interface cultures. The apical medium was subsequently removed, whereby the apical side of the confluent cell layer was exposed to air and an air-liquid interface culture established (Figure 1). To render the HAE-culturing system amenable to genetic modifications, a lentiviral transductionmethodology had to be developed. During the establishment of differentiated HAE-cultures, the primary cells can be transduced at three different stages:
a. after their isolation;
b. in an undifferentiated state during liquid-liquid interface culturing;
c. in a differentiated state during air-liquid interface culturing.
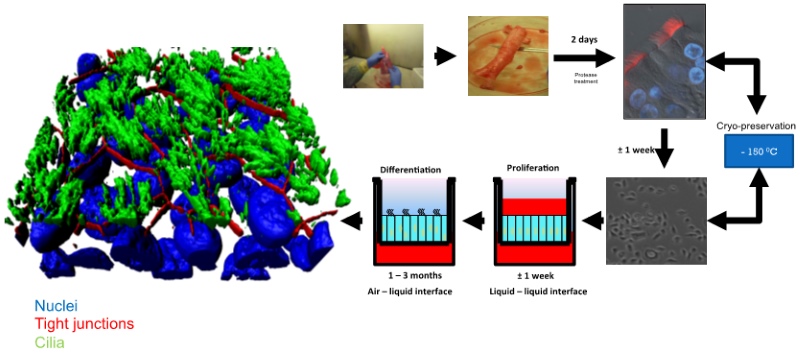
Figure 1: Generation of HAE-cultures: We have assessed the transduction efficacies at these different stages. In suspension, undifferentiated epithelial cells were observed to be most efficiently transduced and expanded as conventional cultures. At a rather low multiplicity of infection (MOI) of 5, we were able to detect transgene expression in 5-30 % of the cells. Importantly, although the cells were transduced during the undifferentiated stage, the transduced cells (i.e. those expressing the transgene) can undergo full differentiation into a pseudostratified epithelial layer (Figure 2). We used a number of marker proteins (Mucin1, EpCAM, NGFR, Tubulin) to assess the cellular composition of cell types that are represented within the epithelium (Figure 2, A&B). These analyses revealed that the cellular compositions are similar between different donors, when different MOIs are used. Moreover, the cellular composition of transgene-expressing cells remained indistinguishable from the non-transduced HAE-cultures.
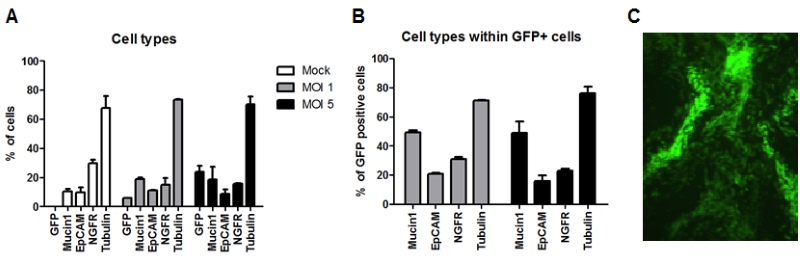
Figure 2: Analysis of transduced and fully-differentiated HAE-cultures: The cellular composition of the HAE-cultures was assessed by flow cytometry (A, B); and transgene expression (GFP) by the transduced and the fully-differentiated HAE-cultures was evaluated by fluorescence microscopy (C).
Increasing the passage number of undifferentiated cells using the Rho-kinase inhibitor Y-27632
In order to obtain epithelial cultures containing (almost) exclusively transduced cells, we employ MACS-sorting in combination with the cell-surface expression of the Cherry-Picker protein. However, one limitation of this approach is that following the initial isolation of the primary human epithelial cells, only 2-3 passages are possible before the cells lose their ability to re-differentiate and to form intact pseudostratified epithelial layers. Since the expression of the Cherry-Picker protein and the cell-sorting process require at least 1-2 passages we established the use of the Rho-kinase inhibitor Y-27632, that has been described to prolong the basal cell phenotype of primary epithelial cells in culture. Using this approach, we were indeed able to propagate primary epithelial cells in vitro for an extended period of time. Most importantly, Y-27632-treated cells were still capable of fully differentiating into pseudostratified epithelial layers in late passages, whereas non-treated cells failed to form fully-differentiated layers after passage 3 (Figure 3). This approach represents a significant improvement, since it is now possible - without any passage or time restriction - to assess the expression of the selection gene (e.g., the Cherry-Picker gene), to select for transduced cells, and even to extend the period of proliferation of the transduced cells to obtain larger quantities (e.g., for storage).
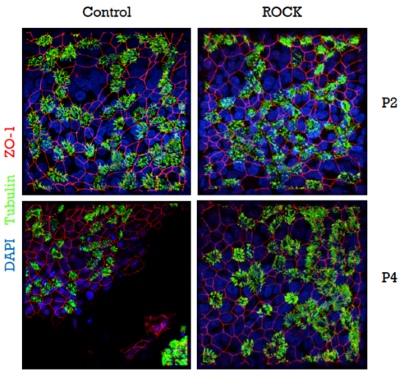
Figure 3: Y-27632-treated cells retain the ability to form intact and fully-differentiated HAE-cultures after passage 3.
Conclusions and Relevance for 3R
The availability of an HAE-culture system that permits to efficiently inactivate the expression of particular genes or the expression of genes of interest in trans renders possible the replacement of many animal experiments that are based on the use of specific transgenic and knock-out mouse strains. An HAE-culturing system that is amenable to the genetic modification of host gene expression is likely to find a wide application in the analysis of host-pathogen interactions appertaining to many human respiratory pathogens.
In addition, the emergence of the MERS-coronavirus during this project afforded us the opportunity of demonstrating the usefulness of primary HAE-cultures for experimental in vitro studies on newly emerging respiratory pathogens. The HAE-culture system could be used to demonstrate the growth kinetics and the cellular tropism of the MERS-coronavirus (1) and to identify its cellular receptor on differentiated human lung epithelia (2).
References
1. Stalin Raj, V., Mou,H., Smits, S.L., Dekkers, D.H.W., Müller, M.A., Dijkman, R., Muth, D., Demmers, J.A.A., Zaki, A., Fouchier, R.A.M., Thiel, V., Drosten, C., Rottier, P.J.M., Osterhaus, A.D.M.E., Bosch, B.J., and Haagmans, B.L. Dipeptidyl peptidase 4 is a functional receptor for the emerging human coronavirus-EMC. 2013. Nature 495:251-254.
2. Kindler, E., Jónsdóttir, H., Muth, D., Hamming, O., Hartmann, R., Rodriguez, R., Geffers, R., Fouchier, R.A.M., Drosten, C., Müller, M.A., Dijkman, R., and Thiel, V. Efficient replication of the novel human betacoronavirus EMC on primary human epithelium highlights its zoonotic potential. 2013. mBio 4(1):doi:10.1128/mBio.00611-12.
| Last modified 2018/10/12 |