
3R-INFO-BULLETIN 5 - August 1995
| | Dr. Monique Vogel, senior investigator in this 3R-project.  With a strong molecular biology background and a new job at the Institute of Immunology and Allergology, Inselspital, Bern, it was clear for her in 1990 that themes such as "repertoire cloning" and "phage display libraries" were very attractive. She devoted all her time to establish these methods and to prove that antibodies can be generated without animal experimentation in the frame of a grant from the Foundation Research 3R. With a strong molecular biology background and a new job at the Institute of Immunology and Allergology, Inselspital, Bern, it was clear for her in 1990 that themes such as "repertoire cloning" and "phage display libraries" were very attractive. She devoted all her time to establish these methods and to prove that antibodies can be generated without animal experimentation in the frame of a grant from the Foundation Research 3R.
Her endeavour as also acknowledged by a prize from the Swiss Society for Animal Research (SGV) in 1994. Since then she co-authored several publications on the isolation of protective antibodies against bacterial toxins, autoantibodies against rare serum proteins or frequent cell surface receptors, and antibodies against red blood cells that may be in therapy in future.
Contact address:
Prof. B M Stadler
Institute of Immunology and Allergology
Inselspital, CH-3011 Bern Phone: 031-632 3521
Fax: 031-381 5735
E-mail: stadler@insel.unibe.ch |
Human recombinant antibodies
Valuable research tools
How do you make an infinite number of different molecules out of a finite number of building blocks? This is one of the tasks of the immune system to construct antibodies that can recognize every possible foreign micro-organism that might enter the body, and thus mediate an immune response against the intruder. The immune system solves this problem by combining several highly variable structural elements (similar to a locksmith combining bolts and pins to make a unique lock) with a few standard elements ("lock") to make approx. 109 variations, each of which reacts with unique specificity and sensitivity to a particular antigen ("key").
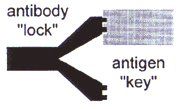 Because of these highly specific "lock and key" reactions, antibodies are invaluable tools in medical research and for therapy. However, until recently it has been necessary to use substantial numbers of animals, challenging them with the antigen and collecting their serum, in order to produce sufficient quantities of antibodies of the desired specificity.
Because of these highly specific "lock and key" reactions, antibodies are invaluable tools in medical research and for therapy. However, until recently it has been necessary to use substantial numbers of animals, challenging them with the antigen and collecting their serum, in order to produce sufficient quantities of antibodies of the desired specificity.
Immune antibody libraries
The research group under Prof. Beda M. Stadler at the Institute of Immunology and Allergology, University of Bern, has chosen a different approach: in a project supported by the Foundation Research 3R, they are using recombinant DNA techniques to try to make the use of animals in obtaining antibodies obsolete [1 ,2]. The researchers take advantage of the antibodies' genetic structural elements in their work and make use of such powerful molecular biological techniques as the polymerase chain reaction.
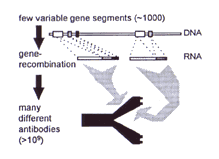
Known regions at either end of the antibody genes were used as primers for the polymerase chain reaction, and the antibody genes were thus selectively amplified to generate an "immune library" from an immunized human blood donor. Genes which encoded for antibodies that were circulating in the blood could be isolated with this method. However, it was not possible to find antibody genes specific for antigens which the donor had never encountered. Thus with this technique [3], just as in the case of the animals, it would be necessary to immunize the donors in order to create recombinant antibodies!
Synthetic libraries
Other research groups have taken a radically different approach, but one that also takes advantage of the standard and variable structural elements of antibodies: they have synthesized their own unique "locks" by generating random variable sequences artificially [4]. From such "synthetic" antibody libraries numerous antibody specifities have been isolated including antibodies against HIV, demonstrating that it may even become obsolete to immunize an individual.
"Panning for gold"
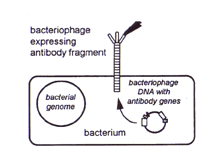
Thus naive (from nonimmunized donors) or immune libraries consisting of ~108, or synthetic libraries consisting of ~1011 different antibodies already exist [1-4]. But, how does the immunologist find the one antibody with the relevant specificity amongst the vast collection? This represents a task like finding a needle in the haystack! The answer lies in another technical breakthrough, namely the use of bacteriophages [3]. These are viruses that can be propagated in bacterial hosts. The genes encoding for the variable region of each antibody are inserted into the DNA of a phage in such a way that the translated antibody protein is expressed on the phage's outer surface. If the antibody is functional, then the phage can be regarded as one large "pseudoantibody".
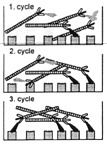 If the library of antibody-carrying phages is exposed to the antigen one is interested in, those which interact can be filtered from those that don't and used for the infection of bacteria. Although the number of phages carrying the desired antibody is small at the start, repeated cycles of antigen exposure and growing allow its isolation in a process that is called "panning", as it resembles the old gold washing procedure.
If the library of antibody-carrying phages is exposed to the antigen one is interested in, those which interact can be filtered from those that don't and used for the infection of bacteria. Although the number of phages carrying the desired antibody is small at the start, repeated cycles of antigen exposure and growing allow its isolation in a process that is called "panning", as it resembles the old gold washing procedure.
Medical applications of recombinant antibodies
Dr. Vogel, the senior investigator in this 3R supported project, has used this technique to isolate autoantibodies against human IgE [1] a molecule associated with allergies or against Tetanus toxin [2]. In the future, such recombinant antibodies may form the basis for the treatment of autoimmune diseases. But there remains a long way to go; at present, the molecules are only fragments containing the variable regions of the antibody. For human therapy the entire antibody molecule, including the constant regions, will have to be constructed and inserted into a suitable vector to be grown in large quantities in a fermentor.
The methods described above are considerably more sophisticated than the standard method of injecting a rabbit with antigen and isolating its antibodies. They also require a certain familiarity with molecular biology techniques. Nevertheless it is hoped that these synthetic antibody libraries will provide a universal tool that will truly make the use of animals for producing antibodies obsolete in the future. It remains to be seen how quickly this technological "revolution" will come about.
References
- Vogel, M., Miescher S., Biaggi, C., Stadler, B. M.
Human anti IgE antibodies by repertoire cloning. Eur. J. Immunol. 24, 1200-1207, 1994. - Lang, A. B., Vogel, M., Viret, J. F., and Stadler B. M.
Polyclonal preparations of anti-Tetanus toxoid antibodies derived from a combinatorial library confer protection. Biotechnology 13, 683-685, 1995. - Nissim, A., Hoogenboom, H. R., Tomlinson, I. M., et al.
Antibody fragments from a single pot phage display library as immunochemical reagents. EMBO J. 13, 692-698, 1994. - Griffiths, A.D., Williams, S.C., Hartley, O., et al. Isolation of high affinity human antibodies directly from large synthetic repertoires. EMBO J. 13, 3245-3260, 1994
With a strong molecular biology background and a new job at the Institute of Immunology and Allergology, Inselspital, Bern, it was clear for her in 1990 that themes such as "repertoire cloning" and "phage display libraries" were very attractive. She devoted all her time to establish these methods and to prove that antibodies can be generated without animal experimentation in the frame of a grant from the Foundation Research 3R.
