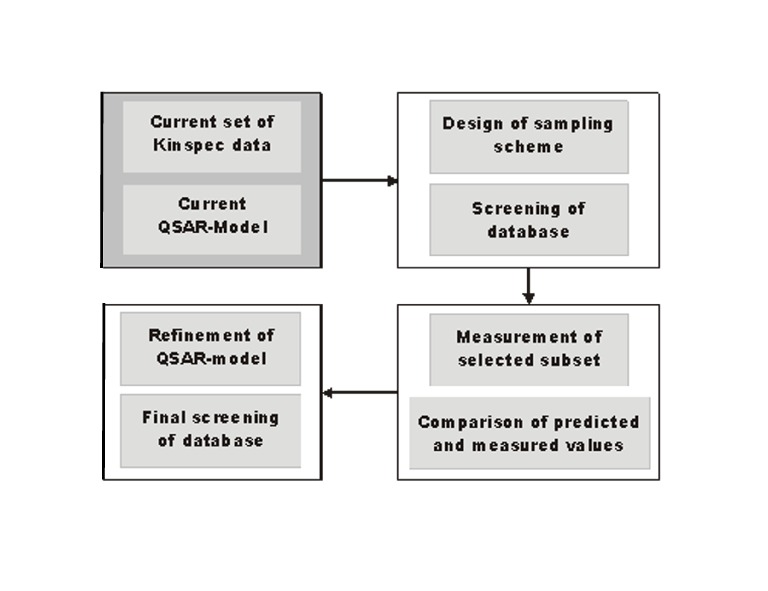
Figure 1: Main steps of research plan. The dark grey box indicates the status at the start of the project.

Beate Escher and Simon Spycher
Eidg. Anstalt für Wasserversorgung, Abwasserreinigung und Gewässerschutz (EAWAG), Abteilung Umwelttoxikologie, 8600 Dübendorf, Switzerland
escher@eawag.ch
Keywords: bacteria; qsar; toxicology; in silico; reduction; replacement
Duration: 1 year Project Completion: 2007
Background and Aim
The pending implementation of the European chemicals regulation REACH (Registration,Evaluation and Authorisation of Chemicals) will have a significant impact on additional testing needs. One approach to potentially reduce the cost and number of animal tests required is the application of quantitative structure-activity relationships (QSAR) as a tool to prioritize further testing. However, in the field of ecotoxicology there are still large gaps for a full regulatory acceptance of QSARs, of which some are difficult to assess in terms of predictive power and often lack transparency.
Over the last years we developed an experimental in vitro test, the Kinspec system (bacterial test), which allows the determination of nonspecific toxic effects in membranes (Escher and Schwarzenbach, 2002). Kinspec covers amongst others the two nonspecific modes of action uncoupling (of oxidative and photo-phosphorylation) and baseline toxicity. One of the advantages of Kinspec is that it allowed new insights into the mechanisms of uncoupling (Escher et al., 1999). These insights lead to the development of a new QSAR model with a strong mechanistic basis and only three physico-chemical descriptors (Spycher et al., 2005).
The proposed project is based on the complementary use of experimental and theoretical work. In the initial phase the existing QSAR-model is used to design a diverse library of potential uncouplers. Subsequently, the compounds in this library will be measured with Kinspec and the results will be compared to the initial predictions of the QSAR model. In a final step a large database of compounds shall be prioritized for further testing. Therefore, this project can be seen as a pilot project for other specific modes of action in ecotoxicologically relevant endpoints to be tested under REACH.
Method and Results
In the Kinspec test system, toxic effects are quantified by changes in membrane potential of chromatophores extracted from the photosynthetic bacterium Rhodobacter sphaeroides. The effect concentrations, EC are derived from the concentration effect curves as described in (Escher and Schwarzenbach, 2002). The distinction of baseline toxicants and uncouplers is based on the calculation of toxic ratios (TR). Compounds with high TRs have a more specific mode of action than baseline toxicity alone. In case of the Kinspec test system, all compounds that accelerate the decay of the membrane potential at concentrations lower than those needed for baseline toxicity must imperatively be charge transporters and if they are additionally weak organic acids, it is clear that they are protonophoric uncouplers.
A set of 21 phenolic uncouplers was used to derive the present QSAR-model. The intrinsic activity model (based on concentrations in the membrane instead of aqueous concentrations) is based on three descriptors: free energies of solvation of the phenolic anion, ∆Gsolv, an empirical parameter for dimer formation and the pKa as a measure for speciation. The two descriptors ∆Gsolv and pKa are independent of (the) chemical class. The dimer formation descriptor is currently limited to phenolic compounds. thus, in a first step a way to generalize this descriptor will be developed. Then a first subset of Kinspec measurements of non phenolic compounds will be used to test if this generalization is successful. The focus of the QSAR-model is to achieve a high robustness, i.e. an ability to generalize to a large group of diverse chemicals. After these first steps the milestones illustrated in Figure 1 will be taken.
The possibility to test an already established model with completely new data is rare in QSAR-modeling and is a remarkable feature of this project. It will give an unbiased picture of the predictive power of the QSAR-model and its potential to screen large databases of thousands of compounds.
Conclusions and Relevance for 3R
From a QSAR perspective the proposed model has two features which make it suitable for regulatory use. The number of descriptors is low and they have a physico-chemical meaning which makes the model transparent, a precondition to legitimize regulatory decisions based on QSAR predictions. The second feature is that it has the potential to be valid not only for one chemical class, but to all compounds acting according to the mode of action of uncoupling and this is still rarely the case in toxicity QSAR. Thus, the proposed project will help priority setting for testing within the framework of the future regulation of chemicals and therefore will save test animals.
References
Escher, B. I., Hunziker, R. P., Schwarzenbach, R. P., Westall, J. C. (1999) Kinetic Model To Describe the Intrinsic Uncoupling Activity of Substituted Phenols in Energy Transducing Membranes. Environ. Sci. Technol., 33, 560-570.
Escher, B., Schwarzenbach, R. P. (2002) Mechanistic studies on baseline toxicity and uncoupling as a basis for modeling internal lethal concentrations in aquatic organisms. Aquat. Sci., 64, 20-35.
Spycher, S., Escher, B., Gasteiger, J. (2005) A QSAR model for the intrinsic activity of uncouplers of oxidative phosphorylation (submitted to Chem. Res. Toxicol.).
Fenner, K., Canonica,S., Escher B. I., Gasser, L.; Spycher, S., Tülp, H. C., (2006) Developing methods to predict chemicaal fate and effect endpoints for use within REACH. Chimia 60/10, 683-690.
Spycher, S., Smejtek, P., Netzeva, T. I., Escher, B. I. (2008), "Towards a class independent QSAR model for uncouplers of oxidative phosphorylation", Chem. Res. Toxicol. (in press).
Figures

Figure 1: Main steps of research plan. The dark grey box indicates the status at the start of the project.