Figure 1

Regine Landmann-Suter
Division of Infectious Diseases, Deüpartment of Research, University Hospital, 4031 Basel, Switzerland
regine.landmann@unibas.ch
Keywords: mice; transgenic mice; kupffer cells; liver; phagocytosis; inflammation; septic shock models; cell cultures: organ-specific; reduction
Duration: 1 year Project Completion: 2002
Background and Aim
Hepatic macrophages (Kupffer cells) are an important cell group because they offer the first line of defence in sepsis. These cells are difficult to study, however, because they survive and maintain their function only for a short time in vitro. Therefore, we established an immortal Kupffer cell line, which would allow the molecular analysis of Kupffer cell function, in particular the study of the interaction between Kupffer cells and bacteria, in vitro
Method and Results
Kupffer cells were isolated from H-2Kb-tsA58 transgenic mice. Four cell lines were cultured at 33ºC in a medium containing interferon-gamma and the supernatants from cell lines of hepatocyte and endothelial cell cultures. The line KC13 was cloned (KC13-2). It grew stably at 37oC without interferon-gamma Its phenotype and function were further studied.
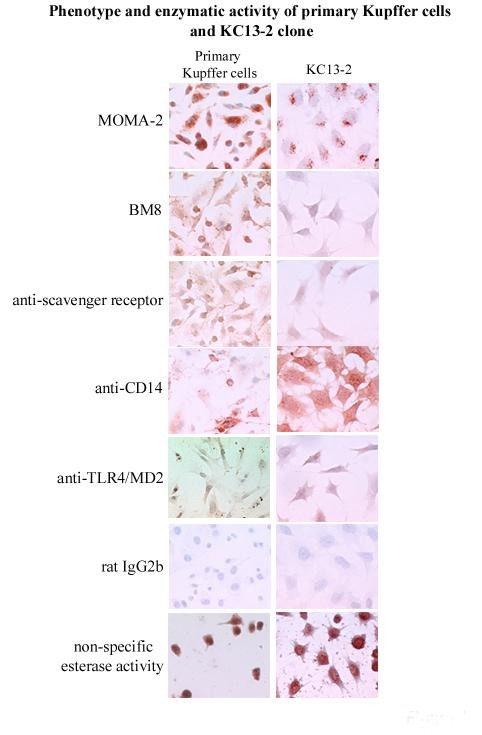
In contrast to primary cells, the cells of the clone were uniform, survived detachment and could therefore be analyzed by cytofluorimetry. The cloned cells expressed many enzymes and markers that are typical of native Kupffer cells (non-specific esterase, peroxidase, MOMA-2, BM8, scavenger receptor A, CD14 and Toll-Like-Receptor 4 (TLR4), the antigen-presenting molecules CD40, CD80 and CD1d, and endocytosed Dextran-FITC. It lacked complement, Fc-receptors, F4/80 marker and the phagosomal coat protein TACO) (Figure 1). The clone exhibited CD14- and TLR4/MD2-independent, plasma-dependent lipopolysaccharide (LPS) binding, E. coli and S. pneumoniae phagocytosis and LPS- and IFN-gamma-induced NO production, but no TNF-alpha, IL-6 or IL-10 release. The clone differed from peritoneal macrophages by the presence of CD1d and the absence of TACO expression and by the fact that LPS-induced functions were independent of CD14 and TLR4.
Conclusions and Relevance for 3R
Conclusions:
We succeeded for the first time in generating a stable, clonal Kupffer cell line.
The large size of the cells, their pattern of surface marker expression and their capacity to clear Gram-negative and -positive bacteria indicate that the clone was derived from the periportal large Kupffer cell subpopulation. Hepatocyte- and endothelial cell-derived factors were required to maintain the differentiated phenotype in the Kupffer cell clone, suggesting that a continuous stimulation is required to keep a differentiation state in this cell type. This property may offer the possibility of identifying which factors modulate gene and protein expression in Kupffer cells and to study interactions with other liver or blood cells.
Relevance:
The set of antigen presenting molecules in the clone allows the consequences of presenting antigens to MHC- and CD1-restricted T cells to be studied. In vitro studies on the consequences of antigen presentation (for example, T cell apoptosis) will also be possible. Moreover, the study of virus or parasite entry into Kupffer cells and the immunological response can be followed in the clone (Figure 2). The clone can be transfected and used to analyze signaling in Kupffer cells after stimulation with bacteria or bacterial products. In summary, the clone allows molecular studies of antiinfective and immune functions of Kupffer cells. It will reduce and replace studies with primary Kupffer cells obtained from mice.
(see also 3R-INFO-BULLETIN Nr. 26
Published updated Version 26/2007 (pdf)
References
Dory Daniel, M Letiembre, H. Echchanaouui, F. Ferracin, J. Pieters, Y. Adachi, S. Akashi, R. Landmann (2003) Generation and functional characterization of a clonal murine periportal Kupffer Cell line form H-2K-tsA58 mice. J Leukocyte Biol. 73, (7) 2003.

Figure 1
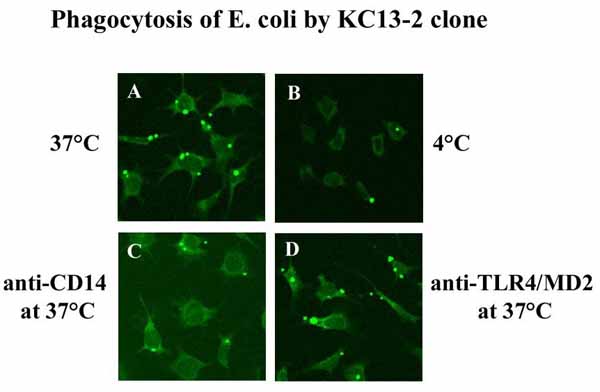
Figure 2