
3R-Project 144-15
Development of in vitro three-dimensional multi-cellular culture models to study the role of heterotypic interactions during colorectal carcinomatous invasive process
Curzio Rüegg, Sarah Cattin
Department of OMI, University of Fribourg, Switzerland
curzio.ruegg@unifr.ch, sarah.cattin@unifr.chKeywords: colorectal cancer; tumour microenvironment; heterotypic interactions; reduction
Duration: 2 years Project Completion: 2018
Background and Aim
Colorectal cancer (CRC) is the third most frequent cause of cancer-related mortality in Europe. Local invasion and distant metastasis are the critical steps that negatively impact patient survival. Although cells of the tumour microenvironment (TME) are known to promote invasion and metastasis, the mechanisms underlying their contribution are only partially understood. A better understanding of how the cells from the TME promote invasion in CRC is crucial for the development of novel anti-invasive therapies.
The study of heterotypic cellular interactions in vivo is difficult, owing to constraints in accessing the tissue and in modulating specific cells or intercellular interactions. In vitro models mimicking the TME would be invaluable in studying the role of individual cells of the TME and their interactions with CRC-cells.
We have developed an in vitro three-dimensional (3D) model to study the interactions between fibroblasts and CRC-cells. This model is now well suited to study the interactions between cancer cells and other cells from the TME, such as endothelial and immune/inflammatory cells. In this project, we aim to improve this 3D in vitro culture model by introducing additional cell types of the TME, and by studying their interactions, with a view to improve our understanding of their role in promoting cancer cell invasion as the first step in the metastatic process. Finally, we will validate the in vivo observations, in order to confirm this test as a robust model to study the heterotypic cellular interactions in the TME.
Method and Results
A 3D in vitro co-culture model consisting of fibroblasts and colorectal cancer cells has already been developed in the past years in our laboratory. This model has been now improved to study the interaction between colorectal cancer cells, fibroblasts and endothelial cells. Cancer cells are embedded in the 3D matrix as spheroid to better mimic a tumour nodule. Spheroids are embedded in the presence of fibroblasts or endothelial cells. As cells are fluorescently-labelled, tumour growth and invasion can be monitored using time-lapse microscopy. 3D cultures can then be treated with cytokines, growth factors, blocking antibodies or pharmacological inhibitors of signalling to determine the key elements that are implicated in cancer cell invasion. To recommend the use of these heterotypic model as an in vitro surrogate of the TME in vivo, it was essential to demonstrate that the in vitro effects of several stromal cells on cancer cells can be reproduced in vivo. To this end, CRC cells were injected subcutaneously or orthotopically into the caecum of immune compromised mice in the absence or presence of fibroblasts or endothelial cells. Progression of the tumour was monitored on the basis of the luciferase activity of the CRC-cells.
Co-cultured fibroblasts promoted SW620 and HCT116 CRC spheroid invasion, and this was prevented by the SRC and FGFR kinase inhibitors, Dasatinib and Erdafitinib respectively. To validate these findings in vivo, SW620 colorectal cancer cells were injected alone or together with fibroblasts orthotopically in the caecum of mice. Co-injection with fibroblasts promoted lung metastasis growth, which was fully reversed by treatment with Dasatinib or Erdafitinib. Co-culture of SW620 or HCT116 colorectal cancer cells spheroids in 3D-Matrigel-matrix in vitro model with endothelial cells suppressed spheroid growth while it had no effect on cancer cell migration or invasion. Consistent with this in vitro effect, co-injected endothelial cells significantly inhibited primary tumour growth in vivo.
From these experiments we concluded that effects on cancer cell invasion and growth induced by co-cultured TME cells and drug treatment in the 3D spheroid model in vitro, are predictive of in vivo effects.
Conclusions and Relevance for 3R
Tumour-host interaction is now emerging to be a critical event in tumour growth and development. However, a study of the TME is a complex undertaking since many cells and factors act simultaneously. A differential evaluation of the underlying mechanisms is thereby extremely difficult in vivo.
The development of this in vitro assay recreates a simplified and well-controlled TME. This 3D spheroid model may be considered as an attractive model to study the effect of heterotypic cellular interactions and drug activities on cancer cells, as animal testing alternative. This model may be adapted and further developed to include different types of cancer and host cells and to investigate additional functions and drugs.
References
- McAllister, S.S., and Weinberg, R.A. (2014). The tumour-induced systemic environment as a critical regulator of cancer progression and metastasis. Nature cell biology 16, 717-727.
- Lorusso, G. & Ruegg, C. The tumour microenvironment and its contribution to tumour evolution toward metastasis. Histochemistry and cell biology 130, 1091-1103, doi:10.1007/s00418-008-0530-8 (2008).
- Lee, G. Y., Kenny, P. A., Lee, E. H. & Bissell, M. J. Three-dimensional culture models of normal and malignant breast epithelial cells. Nature methods 4, 359-365, doi:10.1038/nmeth1015 (2007).
- Kelm, J. M., Timmins, N. E., Brown, C. J., Fussenegger, M. & Nielsen, L. K. Method for generation of homogeneous multicellular tumour spheroids applicable to a wide variety of cell types. Biotechnology and bioengineering 83, 173-180, doi:10.1002/bit.10655 (2003).
- Knuchel, S., Anderle, P., Werfelli, P., Diamantis, E. & Ruegg, C. Fibroblast surface-associated FGF-2 promotes contact-dependent colorectal cancer cell migration and invasion through FGFR-SRC signaling and integrin alphavbeta5-mediated adhesion. Oncotarget (2015).
- Cattin S., Ramont L. & Ruegg C. Characterization and in vivo validation of a three-dimensional multi-cellular culture model to study heterotypic interactions in colorectal cancer cell growth, invasion and metastasis. Front Bioeng Biotechnol. 2018 Jul 17;6:97. doi: 10.3389/fbioe.2018.00097
Figures
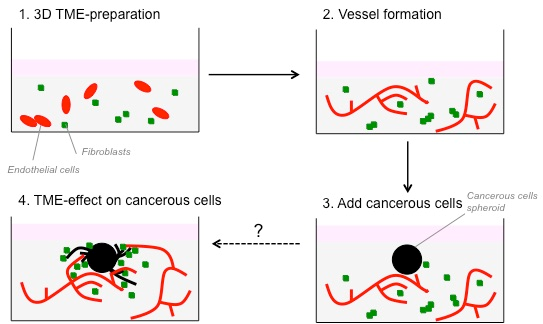
Figure 1: Dasatinib and Erdafitinib inhibitors reduce fibroblasts-induced SW620 cancer cell invasion in vitro under 3D condition. Representative images of SW620-LifeAct-GFP 3D spheroid invasion with and without LifeAct-mCherry labeled fibroblasts in the absence or presence of Dasatinib and Erdafitinib inhibitors after 7 days. Both inhibitors are blocking fibroblasts-induced SW620 3D invasion. Arrows are indicating the invasion area. White bars represent 250 μm.
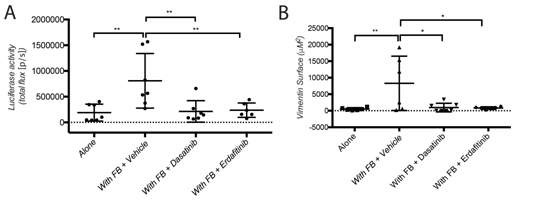
Figure 2: Dasatinib and Erdafitinib suppress fibroblasts-induced metastasis in vivo. (A) Ex-vivo Luciferase activity in the lung of mice orthotopically injected with SW620-A299 cells +/- fibroblasts treated with Dasatinib or Erdafitinib or vehicle only as indicated. (B) Quantification of the metastatic lung surface positive by vimentin by IHC of mice of the same experiment.
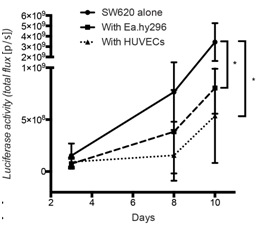
Figure 3: Co-injected endothelial cells reduce SW620 colon cancer growth in vivo.
Quantification of luciferase activity by IBL in mice subcutaneously injected at the indicated conditions over time. Seven mice per group were used. Quantification data represent mean values +/- SD.



